Welcome
Why nlmixr?
The goal of nlmixr, or more accurately nlmixr2, is to support easy and robust nonlinear mixed effects models (NLMEMs) in R.
NLMEMs are used to help identify and explain the relationships between drug exposure, safety, and efficacy and the differences among population subgroups. Most often, they are built using longitudinal PK and pharmacodynamic (PD) data collected during clinical studies. These models characterize the relationships between dose, exposure and biomarker and/or clinical endpoint response over time, variability between individuals and groups, residual variability, and uncertainty.
NLMEM development in the pharmaceutical space is dominated by a small number of proprietary, commercial software tools. Although this kind of approach to software has some advantages, adopting an open-source, open-science paradigm also has benefits - third-party auditing or adjustments are possible, and the precise model-fitting methodology employed can be determined by anyone with the time and energy to review the source code. We see nlmixr2 being especially useful in being able to integrate into the rich R ecosystem, and it is well suited for use in scripted, literate-programming workflows of the kind flourishing in the R ecosystem by means of packages such as knitr and rmarkdown.
The nlmixr2 blog
Survival Analysis with nlmixr2
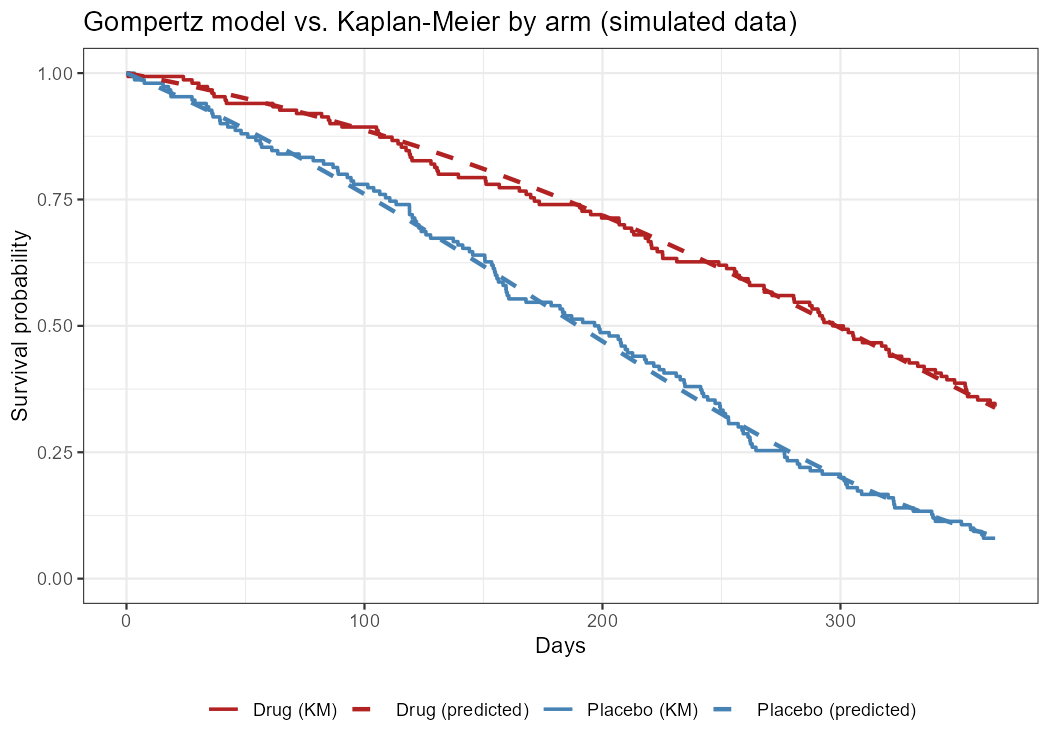
Introduction Survival or time-to-event (TTE) analysis models when something happens, not just whether it happens. The endpoint might be time to tumour progression, time to an adverse event, or time to treatment discontinuation, for instance. In this post we’ll illustrate how to fit parametric TTE models in nlmixr2 using its custom log-likelihood interface. For the purposes of this post, we’re going to simulate a two-arm randomised trial from a known Gompertz model, then recover the parameters with nlmixr2.
Read more